ENPP1 inhibitor with ultralong drug-target residence time as an innate immune checkpoint blockade cancer therapy
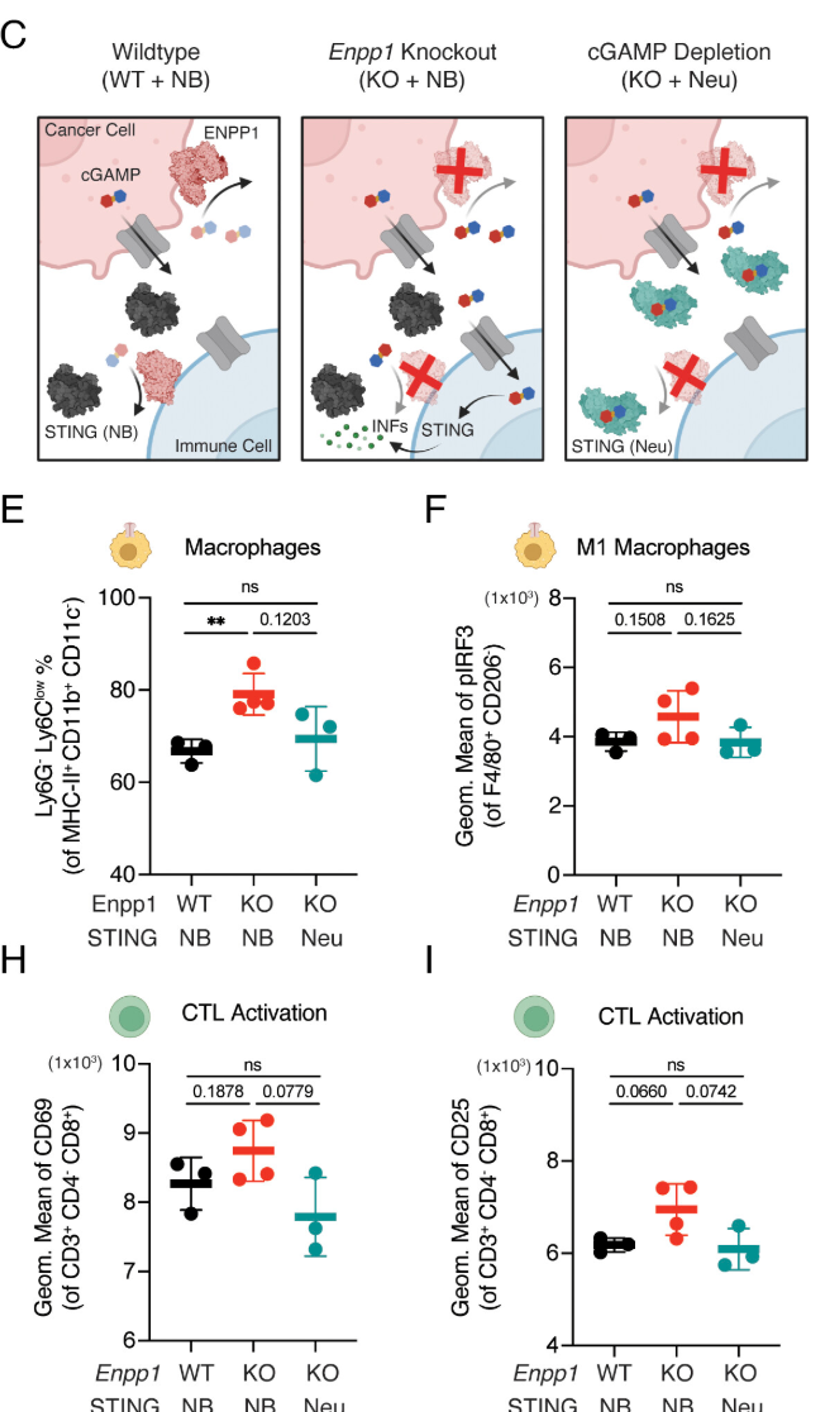
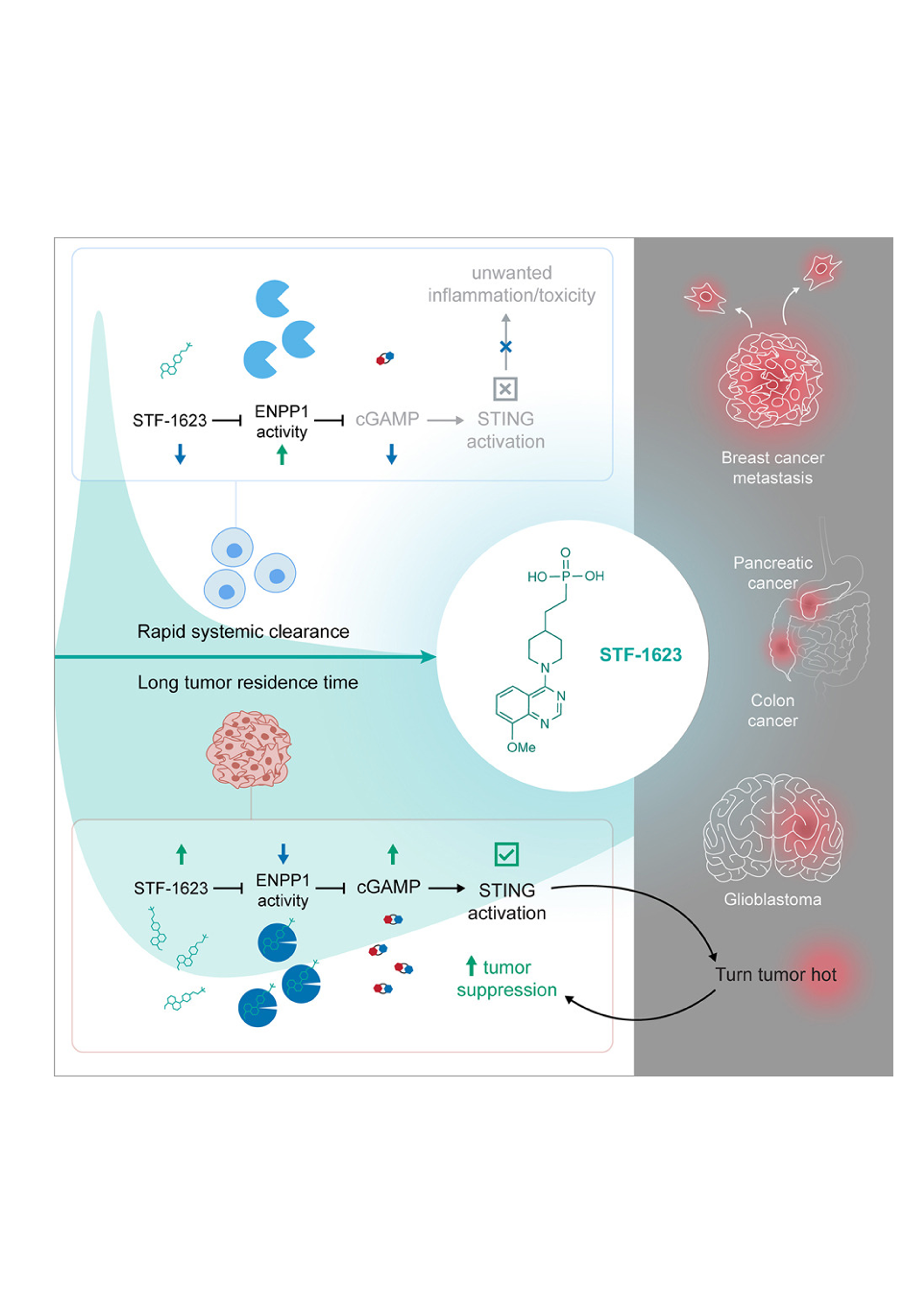
Existing ENPP1 inhibitors have been optimized for prolonged systemic residence time rather than effective target inhibition within tumors. Here, we report the characterization of STF-1623, a highly potent ENPP1 inhibitor with an exceptionally long tumor residence time despite rapid systemic clearance, enabled by its high binding affinity and slow dissociation rate.