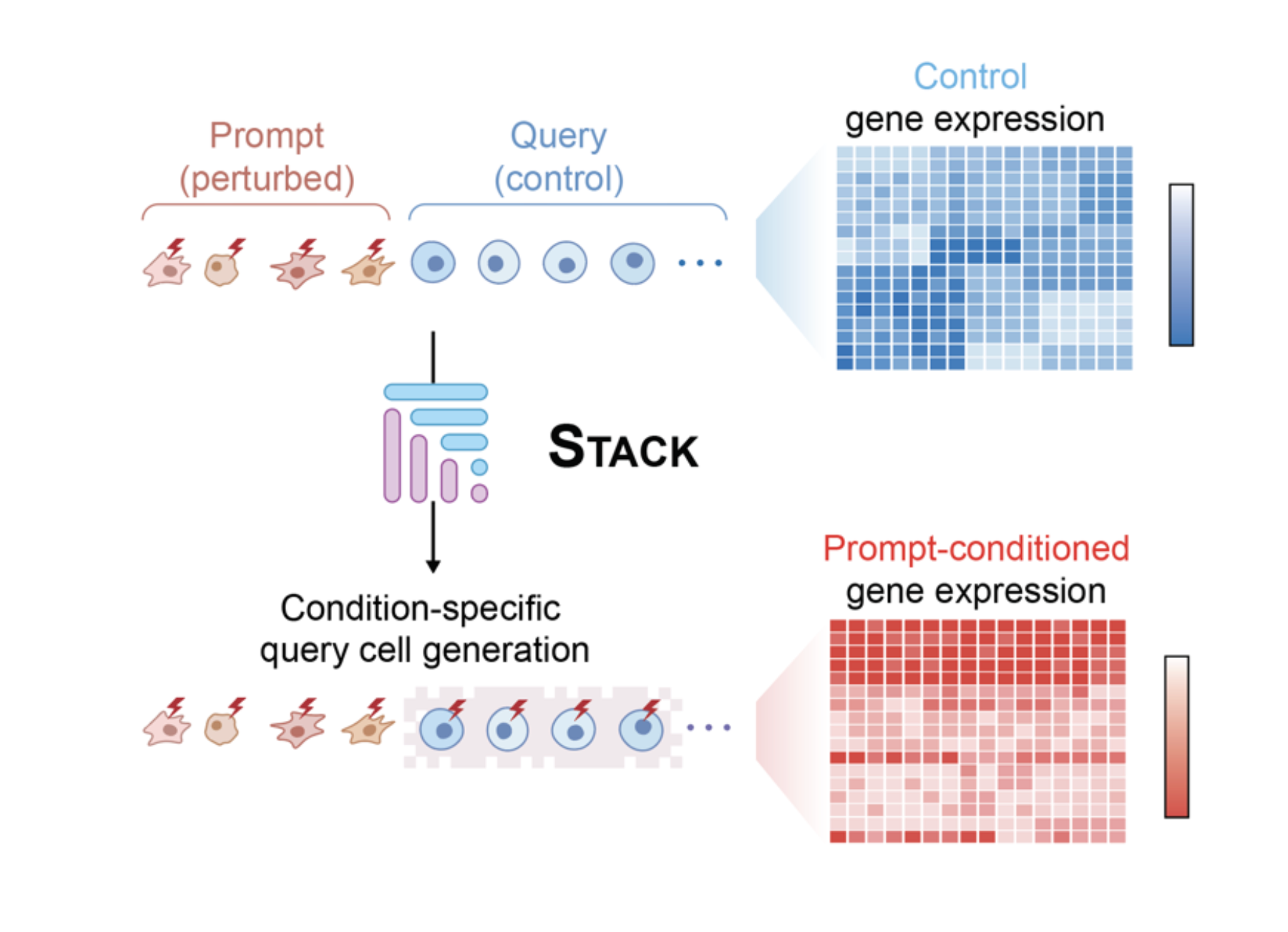
Stack: In-context learning of single-cell biology
Single-cell transcriptomics offers the promise of measuring the diversity of cellular phenotypes across species and diseases. Here, we present Stack, a foundation model trained on 149 million uniformly preprocessed human single cells that leverages tabular attention to generate representations for each cell informed by the cells in its context.