A genome-wide search reveals new antiviral pathways protecting us against COVID-19
While great scientific progress has been made to understand the ongoing COVID-19 pandemic and to develop vaccines that protect against severe outcomes, effective drugs that reduce the likelihood or impact of infection lag behind vaccine efforts and remain a critically important part of our medical response to respiratory virus outbreaks. In order to develop such treatments, a deep molecular understanding of viral infection and interplay with human host cells is essential. A new collaborative effort led by Arc Institute cofounders Patrick Hsu and Silvana Konermann and UC Berkeley virologist Eva Harris has discovered hundreds of human genes and pathways that impact SARS-CoV-2 infection and revealed that airway mucin glycoproteins play a critical role in protecting the lungs from SARS-CoV-2 and other respiratory viruses.
Viruses hijack many cellular functions in order to enter, infect, and replicate in our bodies. To determine which of the over 20,000 human genes are important for a biological process like viral infection, researchers can use CRISPR gene targeting technology to create a pool of millions of human cells, each of which carries a single, unique genetic change somewhere in the genome. Genes can be “knocked out” to be turned off, or “activated” to be turned on. After infecting these cells with a virus and allowing the infection to run its course, researchers can then assess which genetic changes were advantageous or detrimental to cell survival and growth based on which genetic variants are enriched or depleted at the end of the viral challenge.
The new paper from the Hsu and Konermann labs at Arc Institute, and the Harris lab at UC Berkeley, published this week in Nature Genetics, reports the results of such genome-wide CRISPR screens - and notably the first ones conducted in human lung epithelial cells, the major cellular target of SARS-CoV-2 infection in humans. To gain a deeper understanding of viral infection, they not only performed a knockout screen (which would reveal “proviral” genes important for viral replication), but also the first gain-of-function screen for COVID-19 host factors. Using CRISPRa technology originally developed by Konermann, the team hoped that turning on genes that promote cell survival could uniquely reveal “antiviral” genes that prevent viral replication and reveal new therapeutic targets.
CRISPR vs. COVID-19
Conducting genome-wide CRISPR screens with real SARS-CoV-2 virus in high biocontainment wasn’t easy: co-first author Sylvia Sarnik explains, “one of the biggest challenges was scaling up cell culture of the Calu-3 human lung cells that we used – we needed hundreds of millions of them. Thankfully, we found an old paper reporting that certain growth factors could be used to significantly improve the reliable growth of these lung cells, and I think that was really what made a study of this scale possible.” The other co-first author of the work, Scott Biering, agrees: “Virology work is notoriously challenging but working in Biosafety Level 3 (BSL3) is like doing it on ‘hard mode’. Every step takes longer because you have to wipe every inch of every piece of equipment with a decontaminant and you have to be extra focused and careful because you can’t afford any accidents in that environment.”
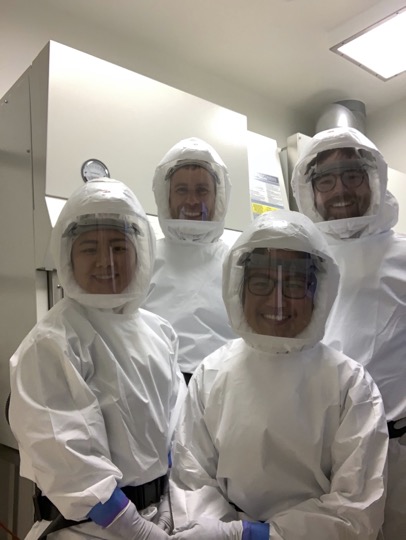

While much of the previous work on SARS-CoV-2 cellular interactions has been conducted on cell types that are easy to grow and manipulate in the lab, they often don’t closely resemble the true cellular targets of SARS-CoV-2 infection in the human airway. “Of course, the more biologically relevant the better, but often the harder it is,” explains Biering. Choosing the right cell type is important because many cells don’t naturally express the major SARS-CoV-2 cell surface receptor, ACE2, or other key factors known to be important in human infections like TMPRSS2, a protease that helps the virus gain entry into our cells. This can force the virus to take other routes into the cell, likely obscuring some important biology at the interface of natural virus-host interactions in the process.
Biering recalls when the team first received the data from their months-long effort: “Patrick emailed me at 2 am saying that our top candidate was ACE2, meaning it worked!” Co-senior author Konermann compared the results to previously published studies, and “it became obvious that we were uncovering the TMPRSS2 pathway, which is the dominant entry route for the virus in real human infections,” but which wasn’t being picked up in other screens that used different cell types. The new paper reports meta-analysis of the findings across several published SARS-CoV-2 screens, demonstrating that the cell type used is the major factor determining differences between the results. The genes that popped up in this study most closely matched genes that are turned on or off in response to infection in COVID-19 patients, providing more evidence for their importance during lung infection.
Combining the results from their knockout and gain-of-function screens allowed the team to come up with a high-confidence list of genes involved in proviral or antiviral pathways. In addition to expected results, like the known viral receptors and entry pathways (ACE2, TMPRSS2, and others) exhibiting proviral effects, they also discovered several new pathways implicated in either promoting or protecting against SARS-CoV-2 infection. Notably, they saw that increased cell growth is detrimental to viral replication, whereas two cellular signaling pathways, NF-kB and interleukin-17, promoted viral replication. “In this paper, we’ve generated so much data – we couldn’t explore everything in mechanistic detail in this paper, but following up on more of those hits would be really exciting for future work,” says Sarnik.
Mucin lung proteins protect against respiratory viruses
A major new finding of the paper is the discovery of a potent antiviral role for airway mucins – glycoproteins that coat the cells in the lung as part of the mucus that lines the airway. “Multiple members of the mucin family lit up in our screens as prominent viral restriction factors, and we found that mucin gene expression also increases as a response to viral infection in patient lungs,” says Hsu.
The team used an enzyme to break down and remove naturally occurring mucins on the cell surface, collaborating with glycoprotein expert Carolyn Bertozzi from Stanford. “We showed that this actually increased the rate of viral infection,” Biering explains, “supporting their protective antiviral role.” The researchers also teamed up with coronavirus and mucin expert Richard Boucher and colleagues from the University of North Carolina, who showed that mice lacking particular mucins were more susceptible to SARS-CoV-2 infection as well. “Working with our collaborators at UNC, we were able to translate our findings from the CRISPR screen into an animal model – and to me, that’s really what makes the paper,” Biering says.
Mechanistically, the team demonstrated that membrane-anchored mucins made it harder for the virus to get into the cells by blocking the attachment of the virus to the cell surface. “By looking at single cell gene expression of lung cells from COVID-19 patients, we found that individual cells with lower levels of mucin expression are more likely to be SARS-CoV-2-positive, again supporting a protective role for mucins in human infections,” says Konermann. This is also consistent with other clinical observations, Biering notes: “Interestingly, with cystic fibrosis patients, who have a huge accumulation of mucus in their lungs, you might expect that they would do worse with COVID-19, but it turns out they actually do better.”
The virus that causes COVID-19 has mutated and given rise to many new variants throughout the course of this pandemic, and each variant is known to behave a bit differently. The researchers wanted to find out whether their mucin findings were broadly applicable across SARS-CoV-2 variants. “In fact, we showed an antiviral role for mucins across all tested variants including the alpha and delta strains,” says Hsu. They also explored whether this might extend to other respiratory viruses, and found that while it was not universal, mucins did have an antiviral impact on several other respiratory viruses including influenza virus PR8, human coronavirus 229E, and human parainfluenza virus PIV3.
With so many different angles to this project, involving CRISPR screens, coronaviruses, human cells, mice, mucins, and glycobiology, this publication is a culmination of a major collaborative effort, and Biering believes that’s why it was so successful: “Things were moving so fast, and you don’t want to compete in the COVID-19 field – we were fortunate to pair up with collaborators who were such experts on mucins and other aspects of the story that really elevated our effort beyond previous work in the field.” Beyond their specific findings on mucin lung proteins, “ambitiously tackling SARS-CoV-2 biology on this massive scale provides a treasure trove of data” that the team hopes other researchers will follow up on, Hsu says. Targeted experiments on new genes and pathways highlighted by this work could lead to future therapeutic targets to boost antiviral defenses or deplete proviral factors to fight against SARS-CoV-2 infection.
